Domains & motifs
Introduction to Domains and Motifs
Protein domains are defined as evolutionary conserved structural or functional units within a protein that have a specific function within the protein. While most proteins consist of at least two domains, however depending on the size and functions of the protein the number of domains varies greatly. The presence and identity of specific domains within a protein can be analyzed to determine the function and evolutionary relationships of proteins [1].
A protein sequence motif is the general term used to refer to specific short conserved regions within a larger sequence [2]. There are several interpretations of the term motif, however all of versions of motifs are defined by short regions of constraint at either the sequence or residue level. In the case of Pendrin a protein primarily characterized by its function in transmembrane transport and localization to the plasma membrane the motifs identified are likely functional motifs, not dependent on invariant residues but highly constrained at the sequence level [2]
A protein sequence motif is the general term used to refer to specific short conserved regions within a larger sequence [2]. There are several interpretations of the term motif, however all of versions of motifs are defined by short regions of constraint at either the sequence or residue level. In the case of Pendrin a protein primarily characterized by its function in transmembrane transport and localization to the plasma membrane the motifs identified are likely functional motifs, not dependent on invariant residues but highly constrained at the sequence level [2]
Domains within Pendrin
STAS Domain
Analysis of the Pendrin protein through the use of domain identification programs SMART, PFAM, and PROSITE show the presence of the STAS domain. STAS is a domain found in the C-terminal cytoplsmic region of a variety of anion transporters. The STAS domain is highly conserved between individuals as well as between species due to the domain's integral role in the function of the protein. The importance of the STAS domain is exemplified by the number of mutations in the SLC26A4 protein resulting in Pendred Syndrome that map to a location that corresponds to the STAS domain location.
SMART |
SMART identified the transmembrane domains of Pendrin, denoted by the blue blocks in the image. The transmembrane regions have multiple alignment regions, evidenced in the image by the multiple blue blocks. The pink bars indicate regions of low sequence complexity. |
PFAM |
PFAM identified three domains within the Pendrin sequence, a sulfate transporter, a sulfate transporter family, and the STAS domain. The green bar represents the sulfate transporter N-terminal domain with a GLY motif localized to the alignment region ranging from 69-152 and characterized with an e-value of 3.5e-29. The sulfate transporter family demonstrated an alignment from 203-481 with an e-value of 1.6e-67. The STAS domain has an alignment from 536-725 and an e-value of 1.6e-42. STAS PFAM accession #: PF01740 Sulfate_transp PFAM accession # PF00916 Sulfate_tra_GLY PFAM accession # PF13792 |
PROSITE |
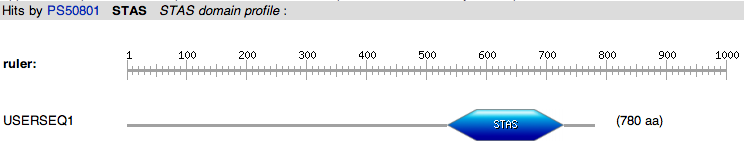
PROSITE identified the presence of the STAS domain and the SLC26A transporters signature. The STAS domain is identified in the alignment region from 535-729 and the SLC26A transporters signature localized to the alignment area of 113-134. STAS PROSITE accession # PS50801 SLC26A transporters signature PROSITE accession # PS01130 |
Motifs
The online MEME program was used to identify the motifs within the pendrin sequences of several species. MEME identified three different motifs, all located within the Sulfate Transporter Family domain, the species specific sequences as well as the logo generated by MEME depicting amino acid variability within the motif are below.
Motif 1
Motif 2
Motif 3
This web page was produced as an assignment for Genetics 564 at UW-Madison Spring 2014
References
[1] Vogel, C., et al. (2004). Structure, function and evolution of multidomain proteins. Current Opinion in Structural Biology. Volume 14. doi: 10.1016/j.sbi.2004.03.011
[2] Bork, P., et al. (1996). Protein sequence motifs. Current Opinion in Structural Biology. Volume 6, Issue 3. doi: 10.1016/S0959-440X(96)80057-1
[3] Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994.
[2] Bork, P., et al. (1996). Protein sequence motifs. Current Opinion in Structural Biology. Volume 6, Issue 3. doi: 10.1016/S0959-440X(96)80057-1
[3] Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994.