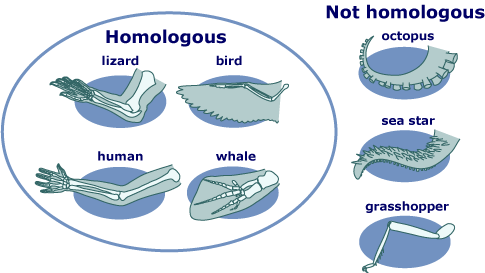
In biological sciences the term homology has multiple definitions depending on the application. Homology defined according to a broad biology approach refers to shared ancestry specific structures between two different species. In genetics, homology is defined as similarity of a region--in the sequence of amino acids comprising a protein or of nucleotides in DNA--between individuals of different species. Homology can be a powerful tool when employed for bioinformatic research. Internet databases present researchers the option of using comparative genomics to predict functions of unknown genes based on sequence homology with nucleotide and protein sequences with known functions.
Introduction to gene homology
Understanding of the evolutionary history of organisms and specifically of particular genes is accomplished through analysis of similarities and differences between organism genomes. Knowledge derived from genome comparisons of different species can be integral in understanding not only the function of a gene but also the evolutionary history of genes within and between species.
Homology is the study of genome similarities between species through inheritance from a common ancestor. A homologous gene is a gene, or variation of a gene, that is present in two different species that can be traced to origination in a common ancestor shared by both of the different species. Identification of genes that are highly conserved in a variety of species can not only elucidate the evolutionary development of current species, but also assist in identification of organisms suitable for laboratory research for further study of specific gene function and disfunction.
Homology is the study of genome similarities between species through inheritance from a common ancestor. A homologous gene is a gene, or variation of a gene, that is present in two different species that can be traced to origination in a common ancestor shared by both of the different species. Identification of genes that are highly conserved in a variety of species can not only elucidate the evolutionary development of current species, but also assist in identification of organisms suitable for laboratory research for further study of specific gene function and disfunction.
Finding homologs of SLC26A4
Homologs of the SLC26A4 gene were found using the tools Homologene and BLAST (Basic Local Alignment Search Tool). These programs analyze input sequences (mRNA or DNA) compared to a database of known and predicted sequences and genomes. The program utelizes a series of algorithms and statistics to provide the user with the genes that statistically show the largest percentage of identical sequence as compared to the input gene. The homologs of the SLC26A4 gene were confirmed through a reverse BLAST to confirm similarity statistics.
Human SLC26A4 Gene
Full gene name: solute carrier family 26 (anion exchanger), member 4 (previously known as PDS gene)
Gene length: 57,174 base pairs
Gene location:
Cytogenic location: 7q31
Molecular location on chromosome 7: base pairs 107,660,634 to 107,717,808
Accession number: NM_000441
Gene length: 57,174 base pairs
Gene location:
Cytogenic location: 7q31
Molecular location on chromosome 7: base pairs 107,660,634 to 107,717,808
Accession number: NM_000441
Homologs of SLC26A4
SLC26A4 has homologous genes in many different species. Highlighted are a few species in which the gene homologous to SLC26A4 has been well studied for phylogenetic analysis, which was performed on Homologene. An E-value of 0 indicates nearly exact sequence similarity in an alignment as compared to the sequence found in humans, the larger the E-value, the less similarity.
Alignments of Homologous Genes
Analysis
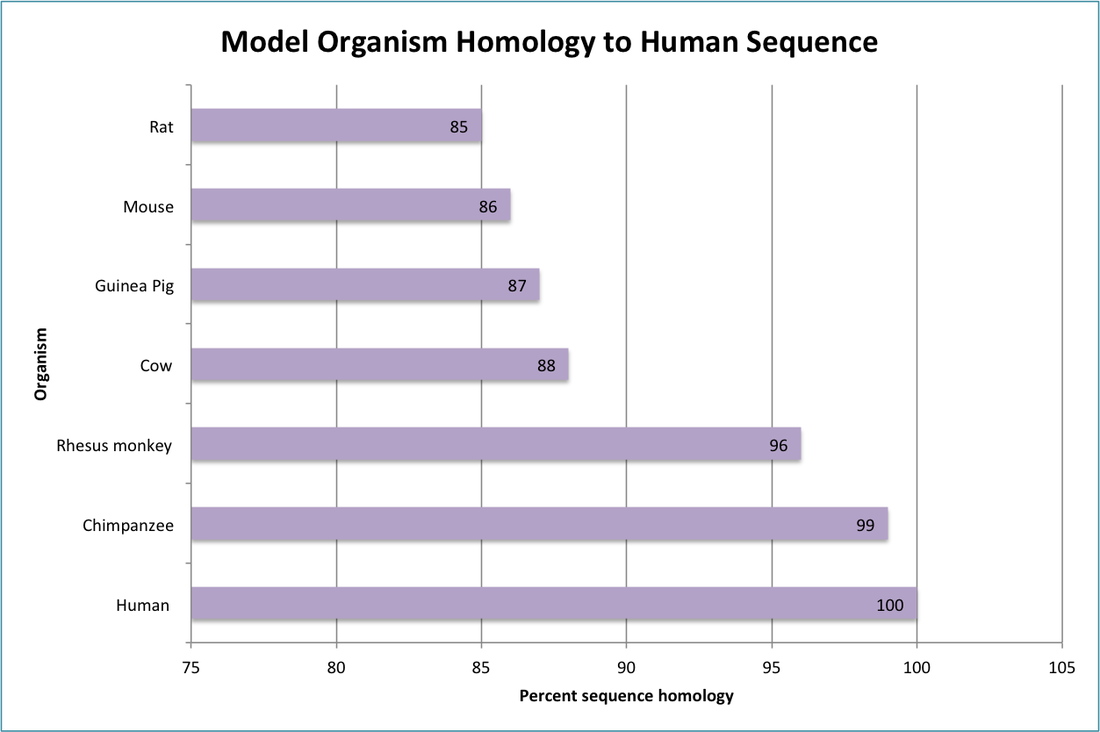
Comparisons of the percentage of identical sequence shared between model organisms and the human sequence of the SLC26A4 gene reveal an expected trend, the model organisms that are close evolutionary relatives of humans, chimpanzees and rhesus monkeys, show the closest gene homology to the human version of the SLC26A4 gene. The remaining organisms (cow, guinea pig, mouse, and rat) also have intuitive homology to the human sequence in accordance with evolutionary relatedness. Homologs to the SLC26A4 gene were limited, likely due to a lack of the phenotype of interest in model organisms that do not have the same inner ear hearing mechanisms as humans. The majority of research on SLC26A4 focuses on elucidating the inner ear mechanism of malformation that causes the congenital deafness associated with Pendred Syndrome, thus making model organisms with highly conserved hearing mechanisms the favored subjects of studies on SLC26A4. There are no homologs to the SLC26A4 gene in non-animal organisms, such as bacteria and plants.
This web page was produced as an assignment for Genetics 564 at UW-Madison Spring 2014